Deploying liquid neural networks to Amazon SageMaker¶
1. Overview¶
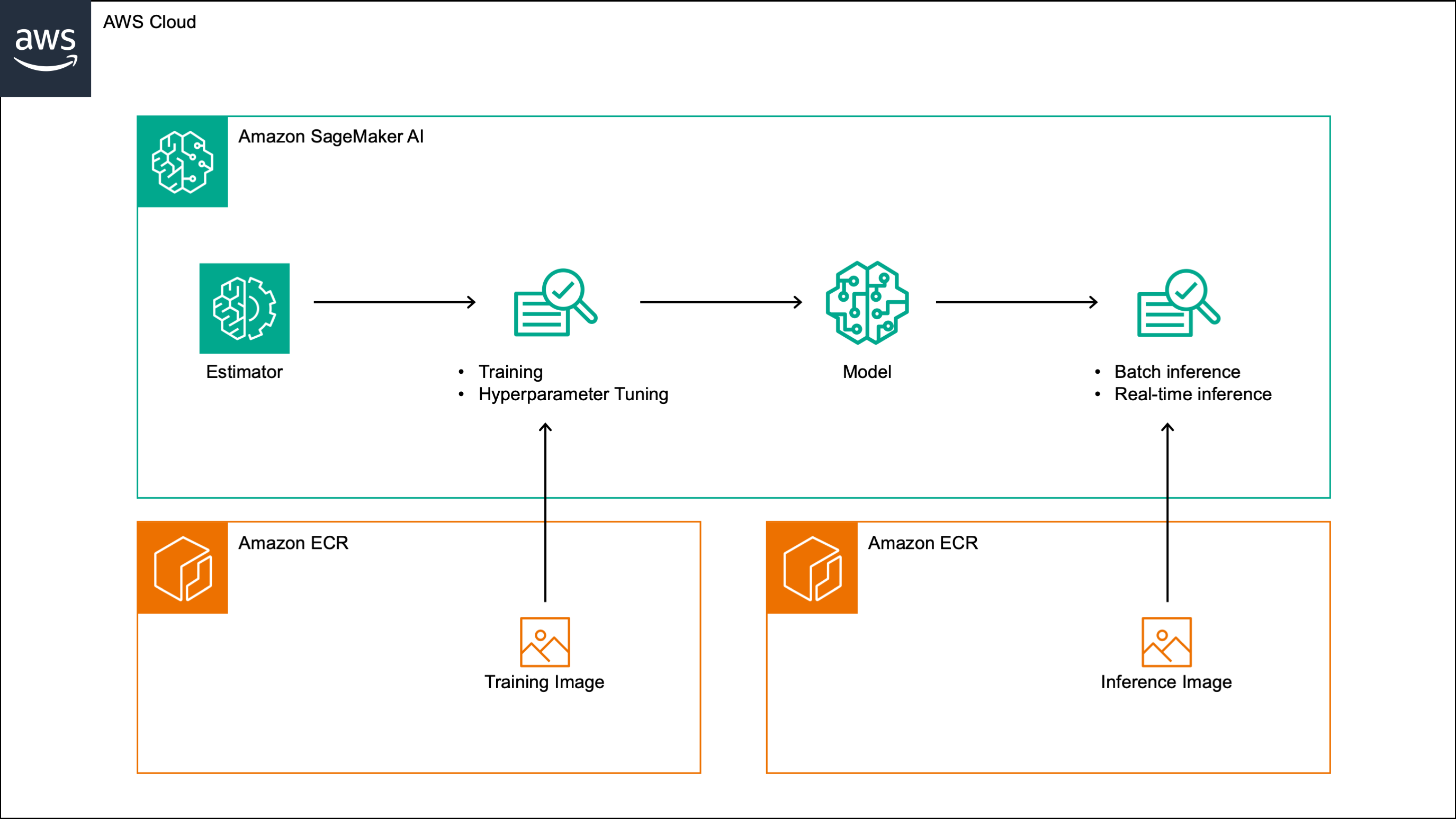
This post walks through a custom Amazon SageMaker algorithm that implements a liquid neural network (LNN) for time series forecasting. The algorithm supports training, incremental training, hyperparameter tuning, batch inference and real-time inference, and can be run on both CPU and GPU instances. It can be used for both univariate and multivariate time series and supports the inclusion of external features. The algorithm minimizes the negative Gaussian log-likelihood during training, and outputs the predicted mean and standard deviation at each future time step, which provide a measure of forecast uncertainty.
The algorithm is built around liquid neural networks (LNNs), a class of continuous-time recurrent neural networks (CT-RNNs) [1, 2] that model the evolution of the hidden state over time as an ordinary differential equation (ODE). LNNs are based on the liquid time constant (LTC) ODE [3], where both the derivative and the time constant of the hidden state are parametrized by a neural network. Like other CT-RNNs, LTCs rely on a numerical ODE solver, which introduces significant computational overhead at both training and inference time. This algorithm implements the closed-form continuous-depth (CfC) variant of LNNs [4], which provides a closed-form approximation to the ODE solution, making it significantly faster than LTCs and other CT-RNNs.
In the rest of this post, we walk through the steps needed to build and test the algorithm. We build the training and inference images by extending the PyTorch 2.1.0 Python 3.10 AWS deep learning containers, and include in both images the original CfC implementation, which is licensed under the Apache License 2.0. We then register the images in a custom Amazon SageMaker algorithm, and test the algorithm on a sample time series dataset in an Amazon SageMaker notebook.
2. Solution¶
The full implementation is available in our GitHub repository. The repository includes the algorithm code, a sample dataset, and a SageMaker notebook that walks through all the steps described in this post. The algorithm code is organized as follows:
amazon_sagemaker_algorithm/
├── modules.py
├── hyperparameters.json
├── training_image/
│ ├── Dockerfile
│ └── model/
│ └── training.py
└── inference_image/
├── Dockerfile
└── inference.py
modules.py contains the original CfC implementation as well as
a custom Model class that reshapes the CfC output into sequences of predicted means and standard deviations,
and stores the data preprocessing parameters alongside the model weights.
hyperparameters.json defines the supported hyperparameters and their specifications.
training_image/ and inference_image/ each contain a Dockerfile and the respective entry
point script, which will be described in detail in Sections 2.1 and 2.2 respectively.
Note
The code below should be run in a SageMaker notebook. The IAM role associated with the notebook should have permissions to read and write to S3 and to push images to ECR.
We start by setting up the environment:
import io
import boto3
import json
import datetime
import sagemaker
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# SageMaker client
sagemaker_client = boto3.client("sagemaker")
# SageMaker session
sagemaker_session = sagemaker.Session()
# SageMaker role
role = sagemaker.get_execution_role()
# S3 bucket
bucket = sagemaker_session.default_bucket()
# EC2 instance
instance_type = "ml.m5.2xlarge"
# AWS account ID
account = sagemaker_session.account_id()
# AWS region
region = "eu-west-1"
2.1 Create the training image¶
We then create the training image. The training_image/ folder contains the Dockerfile
and a model/ subfolder where the training script is stored.
training_image/
├── Dockerfile
└── model/
└── training.py
The script below builds the training image and pushes it to Elastic Container Registry (ECR).
Before building the image, it copies modules.py into the model/ subfolder.
%%bash
algorithm_name=lnn-sagemaker-training
region=eu-west-1
cp amazon_sagemaker_algorithm/modules.py amazon_sagemaker_algorithm/training_image/model/modules.py
cd amazon_sagemaker_algorithm/training_image
account=$(aws sts get-caller-identity --query Account --output text)
fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
aws ecr describe-repositories --repository-names ${algorithm_name} || aws ecr create-repository --repository-name ${algorithm_name}
$(aws ecr get-login --region ${region} --no-include-email)
$(aws ecr get-login --registry-ids 763104351884 --region us-east-1 --no-include-email)
docker build -t ${algorithm_name} . --build-arg REGION=${region}
docker tag ${algorithm_name} ${fullname}
docker push ${fullname}
2.1.1
training.py
training.py reads the hyperparameters from the
command-line arguments, and the input and output paths from the SageMaker environment.
It supports two input data channels:
training (mandatory) and validation (optional). If a model channel is also provided,
the script loads the pre-trained model from the specified directory and continues training it,
otherwise it trains a new model from scratch.
The script infers the column roles from the column names: a column named ts is treated
as time spans, columns starting with x as features, and columns starting with y as targets.
The data is scaled using min-max normalization, with the scaling parameters computed from the training
data and stored in the model checkpoint together with the model and optimizer state dictionaries.
The model is trained using the Adam optimizer with an exponential learning rate decay, minimizing the negative Gaussian log-likelihood. At the end of each epoch, the script logs the mean squared error (MSE) and mean absolute error (MAE) on the training and validation sets, making them available in CloudWatch and as objectives for hyperparameter tuning.
import warnings
warnings.filterwarnings("ignore")
import torch
import argparse
import logging
import tarfile
import os
import sys
import random
from collections import OrderedDict
from sagemaker_training import environment
import pandas as pd
import numpy as np
from modules import Model
logger = logging.getLogger(__name__)
logger.setLevel(logging.INFO)
logger.addHandler(logging.StreamHandler(sys.stdout))
def set_seed(use_cuda):
'''
Fix the random seed, for reproducibility.
'''
random_seed = 0
np.random.seed(random_seed)
random.seed(random_seed)
torch.manual_seed(random_seed)
if use_cuda:
torch.cuda.manual_seed(random_seed)
def get_data(data_dir):
'''
Load the data from the CSV files.
'''
return pd.concat([pd.read_csv(os.path.join(data_dir, f)) for f in os.listdir(data_dir)], axis=0, ignore_index=True)
def get_dataloader(
data,
timespans,
input_length,
output_length,
sequence_stride,
num_outputs,
batch_size,
**kwargs
):
'''
Build the training dataloader.
Note: In this case consecutive sequences can be overlapping
depending on the value of the `sequence_stride` parameter.
'''
t = []
x = []
y = []
for i in range(input_length, len(data) - output_length, sequence_stride):
if timespans is not None:
t.append(timespans[i - input_length: i])
else:
t.append(np.ones(input_length))
x.append(data[i - input_length: i, :])
y.append(data[i: i + output_length, - num_outputs:])
t = np.array(t, dtype=np.float32)
x = np.array(x, dtype=np.float32)
y = np.array(y, dtype=np.float32)
return torch.utils.data.DataLoader(
dataset=torch.utils.data.TensorDataset(
torch.from_numpy(t).float(),
torch.from_numpy(x).float(),
torch.from_numpy(y).float()
),
shuffle=True,
batch_size=batch_size,
**kwargs
)
def get_validation_dataloader(
data,
timespans,
input_length,
output_length,
num_outputs,
batch_size,
**kwargs
):
'''
Build the validation dataloader.
Note: In this case consecutive sequences are non-overlapping.
'''
t = []
x = []
y = []
for i in range(input_length, len(data) - output_length, output_length):
if timespans is not None:
t.append(timespans[i - input_length: i])
else:
t.append(np.ones(input_length))
x.append(data[i - input_length: i, :])
y.append(data[i: i + output_length, - num_outputs:])
t = np.array(t, dtype=np.float32)
x = np.array(x, dtype=np.float32)
y = np.array(y, dtype=np.float32)
return torch.utils.data.DataLoader(
dataset=torch.utils.data.TensorDataset(
torch.from_numpy(t).float(),
torch.from_numpy(x).float(),
torch.from_numpy(y).float()
),
shuffle=False,
batch_size=batch_size,
**kwargs
)
def get_scalers(x):
'''
Calculate the scaling parameters.
'''
min_ = np.nanmin(x, axis=0, keepdims=True)
max_ = np.nanmax(x, axis=0, keepdims=True)
return min_, max_
def mean_squared_error(y_true, y_pred):
'''
Calculate the mean squared error.
'''
return torch.mean(torch.square(y_true - y_pred))
def mean_absolute_error(y_true, y_pred):
'''
Calculate the mean absolute error.
'''
return torch.mean(torch.abs(y_true - y_pred))
def negative_log_likelihood(y, mu, sigma):
'''
Calculate the negative log-likelihood.
'''
return torch.mean(torch.sum(0.5 * torch.tensor(np.log(2 * np.pi)) + 0.5 * torch.log(sigma ** 2) + 0.5 * ((y - mu) ** 2) / (sigma ** 2), dim=-1))
def training_step(
model,
dataloader,
optimizer,
scheduler,
device
):
'''
Run a training step.
'''
model.train()
for t, x, y in dataloader:
t, x, y = t.to(device), x.to(device), y.to(device)
optimizer.zero_grad()
mu, sigma = model(x, t)
loss = negative_log_likelihood(y, mu, sigma)
loss.backward()
optimizer.step()
scheduler.step()
def validation_step(
model,
dataloader,
device
):
'''
Run a validation step.
'''
model.eval()
y_true = []
y_pred = []
for t, x, y in dataloader:
t, x, y = t.to(device), x.to(device), y.to(device)
with torch.no_grad():
yhat, _ = model(x, t)
y_true.append(y.reshape(y.shape[0] * y.shape[1], y.shape[2]))
y_pred.append(yhat.reshape(yhat.shape[0] * yhat.shape[1], yhat.shape[2]))
y_true = torch.cat(y_true, dim=0)
y_pred = torch.cat(y_pred, dim=0)
mse = mean_squared_error(y_true, y_pred).item()
mae = mean_absolute_error(y_true, y_pred).item()
return mse, mae
def fine_tune(args):
'''
Continue training an existing model.
'''
# Extract the environment configuration
use_cuda = torch.cuda.is_available()
use_data_parallel = torch.cuda.device_count() > 1
is_multi_channel = args.test_data_dir is not None
if use_cuda:
device = torch.device("cuda:0")
kwargs = {"num_workers": 1, "pin_memory": True}
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
else:
device = torch.device("cpu")
kwargs = {}
# Load the pre-trained model
print("\n")
print("--------------------------------------")
print("Loading the pre-trained model.")
file = tarfile.open(os.path.join(args.model, 'model.tar.gz'))
file.extractall(args.model)
file.close()
with open(os.path.join(args.model, "model.pth"), "rb") as f:
checkpoint = torch.load(f, map_location="cpu")
params = checkpoint["params"]
model_state_dict = checkpoint["model_state_dict"]
optimizer_state_dict = checkpoint["optimizer_state_dict"]
model = Model(**params)
model.load_state_dict(model_state_dict)
model.to(device)
print(f"Number of parameters: {sum(p.numel() for p in model.parameters())}")
if use_data_parallel:
model = torch.nn.DataParallel(model).to(device)
optimizer = torch.optim.Adam(model.parameters())
optimizer.load_state_dict(optimizer_state_dict)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, args.lr_decay)
print("\n")
print("--------------------------------------")
print("\n")
print("--------------------------------------")
print(f"Initial learning rate: {optimizer_state_dict['param_groups'][0]['lr']}")
print("--------------------------------------")
# Extract the input parameters
timespan_names = params["timespan_names"]
input_names = params["input_names"]
output_names = params["output_names"]
min_ = params["min_"]
max_ = params["max_"]
num_inputs = params["num_inputs"]
num_outputs = params["num_outputs"]
input_length = params["input_length"]
output_length = params["output_length"]
# Load the training data
train_data = get_data(args.train_data_dir)
print("\n")
print("--------------------------------------")
print(f"Training on {train_data.shape[0]} samples.")
print("--------------------------------------")
print("\n")
print("--------------------------------------")
print(f"Timespans: {timespan_names}")
print(f"Features: {input_names}")
print(f"Targets: {output_names}")
print("--------------------------------------")
print("\n")
# Extract the timespans
if len(timespan_names):
train_timespans = train_data.loc[:, timespan_names].values
else:
train_timespans = None
# Reorder the columns
train_data = train_data[input_names + output_names].values
# Scale the data
train_data = (train_data - min_) / (max_ - min_)
if is_multi_channel:
# Load the test data
test_data = get_data(args.test_data_dir)
print("\n")
print("--------------------------------------")
print(f"Validating on {test_data.shape[0]} samples.")
print("--------------------------------------")
print("\n")
# Extract the timespans
if len(timespan_names):
test_timespans = test_data.loc[:, timespan_names].values
else:
test_timespans = None
# Reorder the columns
test_data = test_data[input_names + output_names].values
# Scale the data
test_data = (test_data - min_) / (max_ - min_)
# Build the training dataloader
set_seed(use_cuda)
dataloader = get_dataloader(
train_data,
train_timespans,
input_length,
output_length,
args.sequence_stride,
num_outputs,
args.batch_size,
**kwargs
)
# Build the evaluation dataloaders
training_dataloader = get_validation_dataloader(
train_data,
train_timespans,
input_length,
output_length,
num_outputs,
args.batch_size,
**kwargs
)
if is_multi_channel:
test_dataloader = get_validation_dataloader(
test_data,
test_timespans,
input_length,
output_length,
num_outputs,
args.batch_size,
**kwargs
)
# Train the model
print("\n")
print("--------------------------------------")
print("Training the model.")
set_seed(use_cuda)
for epoch in range(args.epochs):
training_step(model, dataloader, optimizer, scheduler, device)
train_mse, train_mae = validation_step(model, training_dataloader, device)
if is_multi_channel:
valid_mse, valid_mae = validation_step(model, test_dataloader, device)
print(
f'epoch: {format(1 + epoch, ".0f")} '
f'train_mse: {format(train_mse, ",.8f")} '
f'train_mae: {format(train_mae, ",.8f")} '
f'valid_mse: {format(valid_mse, ",.8f")} '
f'valid_mae: {format(valid_mae, ",.8f")}'
)
else:
print(
f'epoch: {format(1 + epoch, ".0f")} '
f'train_mse: {format(train_mse, ",.8f")} '
f'train_mae: {format(train_mae, ",.8f")}'
)
# Score the model
print("\n")
print("--------------------------------------")
print("Scoring the model.")
print("train:mse " + format(train_mse, ',.8f'))
print("train:mae " + format(train_mae, ',.8f'))
if is_multi_channel:
print("valid:mse " + format(valid_mse, ',.8f'))
print("valid:mae " + format(valid_mae, ',.8f'))
print("--------------------------------------")
print("\n")
# Save the model
model.eval()
path = os.path.join(args.model_dir, "model.pth")
model_state_dict = model.state_dict()
optimizer_state_dict = optimizer.state_dict()
if use_data_parallel:
model_state_dict = OrderedDict({k.replace("module.", ""): v for k, v in model_state_dict.items()})
checkpoint = {
"params": params,
"model_state_dict": model_state_dict,
"optimizer_state_dict": optimizer_state_dict,
}
torch.save(checkpoint, path)
def train(args):
'''
Train the model.
'''
# Extract the environment configuration
use_cuda = torch.cuda.is_available()
use_data_parallel = torch.cuda.device_count() > 1
is_multi_channel = args.test_data_dir is not None
if use_cuda:
device = torch.device("cuda:0")
kwargs = {"num_workers": 1, "pin_memory": True}
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
else:
device = torch.device("cpu")
kwargs = {}
# Load the training data
train_data = get_data(args.train_data_dir)
print("\n")
print("--------------------------------------")
print(f"Training on {train_data.shape[0]} samples.")
print("--------------------------------------")
print("\n")
# Extract the variable names
timespan_names = [s for s in train_data.columns if s == "ts"]
input_names = [s for s in train_data.columns if s.startswith("x")]
output_names = [s for s in train_data.columns if s.startswith("y")]
print("\n")
print("--------------------------------------")
print(f"Timespans: {timespan_names}")
print(f"Features: {input_names}")
print(f"Targets: {output_names}")
print("--------------------------------------")
print("\n")
# Calculate the number of variables
num_inputs = len(input_names)
num_outputs = len(output_names)
# Extract the timespans
if len(timespan_names):
train_timespans = train_data.loc[:, timespan_names].values
else:
train_timespans = None
# Reorder the columns
train_data = train_data[input_names + output_names].values
# Calculate the scaling parameters
min_, max_ = get_scalers(train_data)
# Scale the data
train_data = (train_data - min_) / (max_ - min_)
if is_multi_channel:
# Load the test data
test_data = get_data(args.test_data_dir)
print("\n")
print("--------------------------------------")
print(f"Validating on {test_data.shape[0]} samples.")
print("--------------------------------------")
print("\n")
# Extract the timespans
if len(timespan_names):
test_timespans = test_data.loc[:, timespan_names].values
else:
test_timespans = None
# Reorder the columns
test_data = test_data[input_names + output_names].values
# Scale the data
test_data = (test_data - min_) / (max_ - min_)
# Extract the hyperparameters
params = dict(
timespan_names=timespan_names,
input_names=input_names,
output_names=output_names,
min_=min_,
max_=max_,
num_inputs=num_inputs,
num_outputs=num_outputs,
input_length=args.context_length,
output_length=args.prediction_length,
hidden_size=args.hidden_size,
layers=args.backbone_layers,
units=args.backbone_units,
activation=args.backbone_activation,
dropout=args.backbone_dropout,
minimal=int(args.minimal) == 1,
no_gate=int(args.no_gate) == 1,
use_mixed=int(args.use_mixed) == 1,
use_ltc=int(args.use_ltc) == 1,
)
# Build the model
print("\n")
print("--------------------------------------")
print("Building the model.")
set_seed(use_cuda)
model = Model(**params)
model.to(device)
print(f"Number of parameters: {sum(p.numel() for p in model.parameters())}")
if use_data_parallel:
model = torch.nn.DataParallel(model).to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=args.lr)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, args.lr_decay)
print("--------------------------------------")
print("\n")
# Build the training dataloader
set_seed(use_cuda)
dataloader = get_dataloader(
train_data,
train_timespans,
args.context_length,
args.prediction_length,
args.sequence_stride,
num_outputs,
args.batch_size,
**kwargs
)
# Build the evaluation dataloaders
training_dataloader = get_validation_dataloader(
train_data,
train_timespans,
args.context_length,
args.prediction_length,
num_outputs,
args.batch_size,
**kwargs
)
if is_multi_channel:
test_dataloader = get_validation_dataloader(
test_data,
test_timespans,
args.context_length,
args.prediction_length,
num_outputs,
args.batch_size,
**kwargs
)
# Train the model
print("\n")
print("--------------------------------------")
print("Training the model.")
set_seed(use_cuda)
for epoch in range(args.epochs):
training_step(model, dataloader, optimizer, scheduler, device)
train_mse, train_mae = validation_step(model, training_dataloader, device)
if is_multi_channel:
valid_mse, valid_mae = validation_step(model, test_dataloader, device)
print(
f'epoch: {format(1 + epoch, ".0f")}, '
f'train_mse: {format(train_mse, ",.8f")} '
f'train_mae: {format(train_mae, ",.8f")} '
f'valid_mse: {format(valid_mse, ",.8f")} '
f'valid_mae: {format(valid_mae, ",.8f")}'
)
else:
print(
f'epoch: {format(1 + epoch, ".0f")}, '
f'train_mse: {format(train_mse, ",.8f")} '
f'train_mae: {format(train_mae, ",.8f")}'
)
# Score the model
print("\n")
print("--------------------------------------")
print("Scoring the model.")
print(f"train:mse {format(train_mse, ',.8f')}")
print(f"train:mae {format(train_mae, ',.8f')}")
if is_multi_channel:
print(f"valid:mse {format(valid_mse, ',.8f')}")
print(f"valid:mae {format(valid_mae, ',.8f')}")
print("--------------------------------------")
print("\n")
# Save the model
model.eval()
path = os.path.join(args.model_dir, "model.pth")
model_state_dict = model.state_dict()
optimizer_state_dict = optimizer.state_dict()
if use_data_parallel:
model_state_dict = OrderedDict({k.replace("module.", ""): v for k, v in model_state_dict.items()})
checkpoint = {
"params": params,
"model_state_dict": model_state_dict,
"optimizer_state_dict": optimizer_state_dict,
}
torch.save(checkpoint, path)
if __name__ == "__main__":
parser = argparse.ArgumentParser()
parser.add_argument(
"--context-length",
type=int,
)
parser.add_argument(
"--prediction-length",
type=int,
)
parser.add_argument(
"--sequence-stride",
type=int,
)
parser.add_argument(
"--backbone-layers",
type=int,
)
parser.add_argument(
"--backbone-units",
type=int,
)
parser.add_argument(
"--backbone-activation",
type=str,
)
parser.add_argument(
"--backbone-dropout",
type=float,
)
parser.add_argument(
"--hidden-size",
type=int,
)
parser.add_argument(
"--minimal",
type=int,
)
parser.add_argument(
"--no-gate",
type=int,
)
parser.add_argument(
"--use-ltc",
type=int,
)
parser.add_argument(
"--use-mixed",
type=int,
)
parser.add_argument(
"--lr",
type=float,
)
parser.add_argument(
"--lr-decay",
type=float,
)
parser.add_argument(
"--batch-size",
type=int,
)
parser.add_argument(
"--epochs",
type=int,
)
env = environment.Environment()
if len(env.hosts) > 1:
raise ValueError("Distributed training is not supported.")
parser.add_argument(
"--model-dir",
type=str,
default=env.model_dir
)
parser.add_argument(
"--train-data-dir",
type=str,
default=env.channel_input_dirs["training"]
)
parser.add_argument(
"--test-data-dir",
type=str,
default=env.channel_input_dirs["validation"] if "validation" in env.channel_input_dirs else None
)
parser.add_argument(
"--model",
type=str,
default=env.channel_input_dirs["model"] if "model" in env.channel_input_dirs else None
)
args = parser.parse_args()
if args.model is not None:
fine_tune(args)
else:
train(args)
2.1.2
Dockerfile
The Dockerfile extends the AWS PyTorch 2.1.0 GPU training container and copies the contents of
the model/ folder into /opt/ml/code, which is the directory SageMaker expects the training
script to be in. The SAGEMAKER_PROGRAM environment variable tells the container to use
training.py as the entry point.
FROM 763104351884.dkr.ecr.us-east-1.amazonaws.com/pytorch-training:2.1.0-gpu-py310-cu121-ubuntu20.04-sagemaker
ENV PATH="/opt/ml/code:${PATH}"
COPY /model /opt/ml/code
ENV SAGEMAKER_SUBMIT_DIRECTORY /opt/ml/code
ENV SAGEMAKER_PROGRAM training.py
2.2 Create the inference image¶
We now create the inference image. The inference_image/ folder contains the Dockerfile and
inference.py.
inference_image/
├── Dockerfile
└── inference.py
The script below builds the inference image and pushes it to ECR.
As with the training image, modules.py is copied into the inference_image/ folder
before building, since the inference script imports the Model class from it.
%%bash
algorithm_name=lnn-sagemaker-inference
region=eu-west-1
cp amazon_sagemaker_algorithm/modules.py amazon_sagemaker_algorithm/inference_image/modules.py
cd amazon_sagemaker_algorithm/inference_image
account=$(aws sts get-caller-identity --query Account --output text)
fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
aws ecr describe-repositories --repository-names ${algorithm_name} || aws ecr create-repository --repository-name ${algorithm_name}
$(aws ecr get-login --region ${region} --no-include-email)
$(aws ecr get-login --registry-ids 763104351884 --region us-east-1 --no-include-email)
docker build -t ${algorithm_name} . --build-arg REGION=${region}
docker tag ${algorithm_name} ${fullname}
docker push ${fullname}
2.2.1
inference.py
inference.py implements the four handler methods expected by the SageMaker PyTorch serving container:
default_model_fn loads the model checkpoint and restores the model architecture and weights from the
parameters and state dictionary saved during training; default_input_fn deserializes the CSV request
payload into a DataFrame; default_predict_fn runs inference and returns a DataFrame with the predicted
mean and standard deviation for each target column and future time step; default_output_fn serializes
the predictions back to CSV.
Inference runs sequence by sequence, with non-overlapping windows of length prediction-length.
The first context-length rows of the output are set to NaN, since no predictions are generated for
the initial context window. The output also includes prediction-length additional rows beyond the last
input time step, corresponding to the out-of-sample forecast. Predictions are inverse-scaled to the
original data range before being returned.
import warnings
warnings.filterwarnings("ignore")
import os
import io
import pandas as pd
import numpy as np
import torch
from sagemaker_inference import default_inference_handler
from sagemaker_pytorch_serving_container.modules import Model
# Extract the environment configuration
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
use_data_parallel = torch.cuda.device_count() > 1
kwargs = {"num_workers": 1, "pin_memory": True} if torch.cuda.is_available() else {}
class Dataset(torch.utils.data.Dataset):
'''
Define a custom dataset for processing variable length sequences.
'''
def __init__(self, t, x, transform=None):
self.t = t
self.x = x
self.transform = transform
def __len__(self):
return len(self.x)
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.item()
return [
torch.from_numpy(self.t[idx]).float(),
torch.from_numpy(self.x[idx]).float(),
]
def get_test_dataloader(
data,
timespans,
input_length,
output_length,
**kwargs
):
'''
Build the dataloader using the custom dataset.
'''
t = []
x = []
for i in range(input_length, len(data) + output_length, output_length):
if timespans is not None:
t.append(timespans[i - input_length: i])
else:
t.append(np.ones(input_length))
x.append(data[i - input_length: i, :])
return torch.utils.data.DataLoader(
dataset=Dataset(t, x),
shuffle=False,
batch_size=1,
**kwargs
)
class DefaultPytorchInferenceHandler(default_inference_handler.DefaultInferenceHandler):
def default_model_fn(self, model_dir):
'''
Load the model.
'''
with open(os.path.join(model_dir, "model.pth"), "rb") as f:
checkpoint = torch.load(f, map_location="cpu")
params = checkpoint["params"]
state_dict = checkpoint["model_state_dict"]
model = Model(**params)
model.load_state_dict(state_dict)
model.to(device)
if use_data_parallel:
model = torch.nn.DataParallel(model).to(device)
return model
def default_input_fn(self, input_data, content_type):
'''
Load the data.
'''
return pd.read_csv(io.StringIO(input_data)).astype(float)
def default_predict_fn(self, data, model):
'''
Generate the predictions.
'''
# Extract the input parameters
if use_data_parallel:
timespan_names = model.module.timespan_names
input_names = model.module.input_names
output_names = model.module.output_names
min_ = model.module.min_
max_ = model.module.max_
input_length = model.module.input_length
output_length = model.module.output_length
num_inputs = model.module.num_inputs
num_outputs = model.module.num_outputs
else:
timespan_names = model.timespan_names
input_names = model.input_names
output_names = model.output_names
min_ = model.min_
max_ = model.max_
input_length = model.input_length
output_length = model.output_length
num_inputs = model.num_inputs
num_outputs = model.num_outputs
# Extract the timespans
if len(timespan_names):
timespans = data.loc[:, timespan_names].values
else:
timespans = None
# Reorder the columns
data = data[input_names + output_names].values
# Scale the data
data = (data - min_) / (max_ - min_)
# Create the dataloader
dataloader = get_test_dataloader(
data,
timespans,
input_length,
output_length,
**kwargs
)
# Generate the model predictions
mu = torch.from_numpy(np.nan * np.ones((input_length, num_outputs))).float().to(device)
sigma = torch.from_numpy(np.nan * np.ones((input_length, num_outputs))).float().to(device)
for t, x in dataloader:
t, x = t.to(device), x.to(device)
with torch.no_grad():
mu_, sigma_ = model(x, t)
mu = torch.cat([mu, mu_.reshape(mu_.size(0) * mu_.size(1), mu_.size(2))], dim=0)
sigma = torch.cat([sigma, sigma_.reshape(sigma_.size(0) * sigma_.size(1), sigma_.size(2))], dim=0)
# Transform the model predictions back to the original scale
mu = min_[:, - num_outputs:] + (max_[:, - num_outputs:] - min_[:, - num_outputs:]) * mu.detach().cpu().numpy()
sigma = (max_[:, - num_outputs:] - min_[:, - num_outputs:]) * sigma.detach().cpu().numpy()
# Organize the model predictions in a data frame
prediction = {}
for i in range(num_outputs):
prediction[output_names[i] + '_mean'] = mu[:, i]
prediction[output_names[i] + '_std'] = sigma[:, i]
prediction = pd.DataFrame(prediction)
return prediction
def default_output_fn(self, prediction, accept):
'''
Return the predictions.
'''
csv_buffer = io.StringIO()
prediction.to_csv(csv_buffer, index=False)
return csv_buffer.getvalue()
2.2.2
Dockerfile
The Dockerfile extends the AWS PyTorch 2.1.0 GPU inference container.
inference.py replaces the default inference handler by overwriting
default_pytorch_inference_handler.py in the sagemaker_pytorch_serving_container package,
and modules.py is copied into the same directory to make the Model class available at
serving time.
FROM 763104351884.dkr.ecr.us-east-1.amazonaws.com/pytorch-inference:2.1.0-gpu-py310-cu118-ubuntu20.04-sagemaker
COPY inference.py /opt/conda/lib/python3.10/site-packages/sagemaker_pytorch_serving_container/default_pytorch_inference_handler.py
COPY modules.py /opt/conda/lib/python3.10/site-packages/sagemaker_pytorch_serving_container/modules.py
2.3 Create the algorithm¶
With both images pushed to ECR, we register them as a custom SageMaker algorithm.
The algorithm specification has two parts: the TrainingSpecification
and the InferenceSpecification.
The TrainingSpecification defines the training image URI, the supported hyperparameters
(loaded from hyperparameters.json), the metric regex patterns used to extract the MSE and MAE
from the training logs, and the three input channels: training (required, CSV), validation
(optional, CSV) and model (optional, Gzip), the latter being used for incremental training.
It also declares valid_mse and valid_mae as the supported tuning objectives.
The InferenceSpecification defines the inference image URI and the supported instance types
for real-time and batch inference, with CSV as both the input content type and the response MIME type.
# Load the list of supported hyperparameters
with open("amazon_sagemaker_algorithm/hyperparameters.json", "r") as f:
supported_hyperparameters = json.load(f)
# Create the algorithm
response = sagemaker_client.create_algorithm(
AlgorithmName='lnn-sagemaker',
TrainingSpecification={
'TrainingImage': f'{account}.dkr.ecr.{region}.amazonaws.com/lnn-sagemaker-training:latest',
'SupportedHyperParameters': supported_hyperparameters,
'SupportsDistributedTraining': False,
'MetricDefinitions': [
{
'Name': 'train_mse',
'Regex': 'train_mse: ([0-9\.]+)'
},
{
'Name': 'train_mae',
'Regex': 'train_mae: ([0-9\.]+)'
},
{
'Name': 'valid_mse',
'Regex': 'valid_mse: ([0-9\.]+)'
},
{
'Name': 'valid_mae',
'Regex': 'valid_mae: ([0-9\.]+)'
},
],
'TrainingChannels': [
{
'Name': 'training',
'IsRequired': True,
'SupportedContentTypes': [
'text/csv',
],
'SupportedInputModes': [
'File',
]
},
{
'Name': 'validation',
'IsRequired': False,
'SupportedContentTypes': [
'text/csv',
],
'SupportedInputModes': [
'File',
]
},
{
'Name': 'model',
'IsRequired': False,
'SupportedContentTypes': [
'Gzip',
],
'SupportedInputModes': [
'File',
]
},
],
'SupportedTuningJobObjectiveMetrics': [
{
'Type': 'Minimize',
'MetricName': 'valid_mse'
},
{
'Type': 'Minimize',
'MetricName': 'valid_mae'
},
],
'SupportedTrainingInstanceTypes': [
instance_type
],
},
InferenceSpecification={
'Containers': [
{
'Image': f'{account}.dkr.ecr.{region}.amazonaws.com/lnn-sagemaker-inference:latest',
}
],
'SupportedContentTypes': [
'text/csv'
],
'SupportedResponseMIMETypes': [
'text/csv'
],
'SupportedTransformInstanceTypes': [
instance_type
],
'SupportedRealtimeInferenceInstanceTypes': [
instance_type
],
},
CertifyForMarketplace=False,
)
Once registered, the algorithm is identified by its ARN, which is used in all subsequent steps.
# Get the algorithm’s ARN
algorithm_arn = response['AlgorithmArn']
2.4 Test the algorithm¶
2.4.1 Upload the sample data to S3¶
We test the algorithm on the sample dataset included in the GitHub repository.
The dataset includes two target time series (y1, y2), two feature time series (x1, x2) and a time
span column (ts), split into training, validation and test sets of 1000, 500 and 500 samples
respectively.
We start by loading the three dataset splits into DataFrames.
# Load the training data
training_dataset = pd.read_csv("sample_data/train.csv")
# Load the validation data
validation_dataset = pd.read_csv("sample_data/valid.csv")
# Load the test data
test_dataset = pd.read_csv("sample_data/test.csv")
We then upload them to S3 to make them available to the training and inference jobs.
# Upload the training data to S3
training_data = sagemaker_session.upload_string_as_file_body(
body=training_dataset.to_csv(index=False),
bucket=bucket,
key="sample_data/train.csv"
)
# Upload the validation data to S3
validation_data = sagemaker_session.upload_string_as_file_body(
body=validation_dataset.to_csv(index=False),
bucket=bucket,
key="sample_data/valid.csv"
)
# Upload the test data to S3
test_data = sagemaker_session.upload_string_as_file_body(
body=test_dataset.to_csv(index=False),
bucket=bucket,
key="sample_data/test.csv"
)
2.4.2 Run a training job¶
We train the model using AlgorithmEstimator, which takes the algorithm ARN instead of a
container image URI. The hyperparameters define a CfC network with a single backbone layer of
128 units and SiLU activation, a hidden size of 64, no dropout, and the default CfC variant
(minimal=0, no-gate=0, use-ltc=0, use-mixed=0). The context length is set to
200 and the prediction length to 100, meaning the model sees 200 time steps to forecast the
next 100. The sequence stride of 1 means consecutive training windows are shifted by one step,
maximizing the number of training samples. The model is trained for 50 epochs with a batch size
of 64, a learning rate of 0.001 and an exponential decay factor of 0.999. The estimator is
fitted by passing the S3 URIs of the training and validation channels.
# Define the hyperparameters
hyperparameters = {
"context-length": 200,
"prediction-length": 100,
"sequence-stride": 1,
"backbone-layers": 1,
"backbone-units": 128,
"backbone-activation": "silu",
"backbone-dropout": 0,
"hidden-size": 64,
"minimal": 0,
"no-gate": 0,
"use-ltc": 0,
"use-mixed": 0,
"lr": 0.001,
"lr-decay": 0.999,
"batch-size": 64,
"epochs": 50,
}
# Create the estimator
estimator = sagemaker.algorithm.AlgorithmEstimator(
algorithm_arn=algorithm_arn,
base_job_name="lnn-training",
role=role,
instance_count=1,
instance_type=instance_type,
input_mode="File",
sagemaker_session=sagemaker_session,
hyperparameters=hyperparameters,
)
# Run the training job
estimator.fit({"training": training_data, "validation": validation_data})
2.4.3 Run an incremental training job¶
To continue training from the previous checkpoint, we pass the model artifacts from the completed
training job to the model_uri parameter of AlgorithmEstimator. SageMaker makes the
artifacts available to the training container via the model input channel, where the training
script detects them, loads the model and optimizer state, and resumes training. The learning rate
is not reset but picked up from the saved optimizer state, so the decay applied during the first training
job carries over. We continue training the model for another 50 epochs using the same hyperparameters.
# Get the URI of the pre-trained model
model_uri = f's3://{bucket}/{estimator.latest_training_job.name}/output/model.tar.gz'
# Create the estimator
estimator = sagemaker.algorithm.AlgorithmEstimator(
model_uri=model_uri,
algorithm_arn=algorithm_arn,
base_job_name="lnn-fine-tuning",
role=role,
instance_count=1,
instance_type=instance_type,
input_mode="File",
sagemaker_session=sagemaker_session,
hyperparameters=hyperparameters
)
# Run the training job
estimator.fit({"training": training_data, "validation": validation_data})
2.4.4 Run a hyperparameter tuning job¶
We use HyperparameterTuner to search over the architecture and training hyperparameters,
optimizing valid_mae. Before creating the tuner, we detach the pre-trained model from the
estimator so that each tuning trial trains from scratch. The search space covers the number of
backbone layers, backbone units, activation function, dropout rate, hidden size, learning
rate, learning rate decay, batch size and number of epochs. The fixed hyperparameters
context-length, prediction-length, sequence-stride and the CfC variant flags are
kept at the values defined in Section 2.4.2. For illustration purposes we run 4 trials in
parallel with a random search strategy, and retrieve the best hyperparameters at the end.
# Detach the pre-trained model
estimator.model_uri = None
# Define the hyperparameter ranges
hyperparameter_ranges = {
"backbone-layers": sagemaker.parameter.IntegerParameter(1, 3),
"backbone-units": sagemaker.parameter.CategoricalParameter([32, 64, 128, 256]),
"backbone-activation": sagemaker.parameter.CategoricalParameter(["silu", "relu", "tanh", "gelu", "lecun"]),
"backbone-dropout": sagemaker.parameter.ContinuousParameter(0, 0.5),
"hidden-size": sagemaker.parameter.CategoricalParameter([32, 64, 128, 256]),
"lr": sagemaker.parameter.ContinuousParameter(0.0001, 0.01),
"lr-decay": sagemaker.parameter.ContinuousParameter(0.9, 1.0),
"batch-size": sagemaker.parameter.CategoricalParameter([32, 64, 128, 256]),
"epochs": sagemaker.parameter.IntegerParameter(20, 200),
}
# Create the hyperparameter tuner
tuner = sagemaker.tuner.HyperparameterTuner(
estimator=estimator,
base_tuning_job_name="lnn-tuning",
objective_metric_name="valid_mae",
objective_type="Minimize",
hyperparameter_ranges=hyperparameter_ranges,
max_jobs=4,
max_parallel_jobs=4,
random_seed=100,
)
# Run the hyperparameter tuning job
tuner.fit({"training": training_data, "validation": validation_data})
# Get the best hyperparameters
tuning_job_result = sagemaker_session.sagemaker_client.describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.name
)
print("Best hyperparameters:")
tuning_job_result["BestTrainingJob"]["TunedHyperParameters"]
2.4.5 Run a batch transform job¶
We create a transformer from the estimator and run a batch transform job on the test set.
SageMaker passes the input CSV to the inference container, which returns a CSV with four columns:
y1_mean, y1_std, y2_mean and y2_std. The output has 600 rows - 500 matching the
test input plus 100 out-of-sample forecasts - with the first 200 rows set to NaN since no
predictions are generated for the initial context window. The predictions are loaded from S3
and parsed into a DataFrame. We delete the model at the end to avoid unnecessary storage costs.
# Create the transformer
transformer = estimator.transformer(
instance_count=1,
instance_type=instance_type,
)
transformer.base_transform_job_name = "lnn-transform"
# Run the batch transform job.
transformer.transform(
data=test_data,
content_type="text/csv",
)
# Load the predictions from S3
batch_predictions = sagemaker_session.read_s3_file(
bucket=bucket,
key_prefix=f"{transformer.latest_transform_job.name}/test.csv.out"
)
batch_predictions = pd.read_csv(io.StringIO(batch_predictions), dtype=float)
# Delete the model
transformer.delete_model()
2.4.6 Perform real-time inference¶
We deploy the model to a real-time endpoint and invoke it by passing
the test CSV directly in the request body. The response is deserialized into a DataFrame using
PandasDeserializer. The output has the same structure as the batch transform output: four
columns (y1_mean, y1_std, y2_mean, y2_std) with 600 rows, the first 200 of which
are NaN. We delete the model and endpoint configuration at the end to avoid ongoing charges.
# Create the endpoint
predictor = estimator.deploy(
initial_instance_count=1,
instance_type=instance_type,
model_name=f"lnn-model-{datetime.datetime.now().strftime(format='%Y-%m-%d-%H-%M-%S-%f')}",
endpoint_name=f"lnn-endpoint-{datetime.datetime.now().strftime(format='%Y-%m-%d-%H-%M-%S-%f')}",
)
# Invoke the endpoint
response = sagemaker_session.sagemaker_runtime_client.invoke_endpoint(
EndpointName=predictor.endpoint_name,
ContentType="text/csv",
Body=test_dataset.to_csv(index=False)
)
# Extract the predictions from the response
deserializer = sagemaker.base_deserializers.PandasDeserializer(accept="text/csv")
real_time_predictions = deserializer.deserialize(response["Body"], content_type="text/csv")
# Delete the model
predictor.delete_model()
# Delete the endpoint
predictor.delete_endpoint(delete_endpoint_config=True)
References¶
[1] Chow, T.W. and Li, X.D., 2000. Modeling of continuous time dynamical systems with input by recurrent neural networks. IEEE Transactions on Circuits and Systems I: Fundamental Theory and Applications, 47(4), pp.575-578. doi: 10.1109/81.841860.
[2] Funahashi, K.I. and Nakamura, Y., (1993). Approximation of dynamical systems by continuous time recurrent neural networks. Neural networks, 6(6), pp.801-806. doi: 10.1016/S0893-6080(05)80125-X.
[3] Hasani, R., Lechner, M., Amini, A., Rus, D., & Grosu, R. (2021). Liquid time-constant networks. In Proceedings of the AAAI Conference on Artificial Intelligence, 35(9), pp. 7657-7666. doi: 10.1609/aaai.v35i9.16936.
[4] Hasani, R., Lechner, M., Amini, A., Liebenwein, L., Ray, A., Tschaikowski, M., Teschl, G. and Rus, D., (2022). Closed-form continuous-time neural networks. Nature Machine Intelligence, 4(11), pp. 992-1003. doi: 10.1038/s42256-022-00556-7.